|
 |
INTRODUCTION: Submitted for your review is the online report of the experiment, Evidence of Two-Step Recursive Pre-mRNA Splicing In Vitro. Above are links to download the written report and the flash presentation as presented on Febuary 11th, 2005. This experiment examines the post-transcriptional process of messenger RNA splicing. To review, advanced organisms all contain DNA - a long chain consisting of a four letter code that provides the structural information of the organism. In a process called Transcription, the DNA code is converted into a new code comprised of RNA. These shorter segments of code can then be interpreted to create proteins in a process called translation. Proteins are the building blocks for all cells and ultimately of the organism itself.
The human genome project has recently completed the task of sequencing the entire human DNA code. By doing so they have been able to determine areas of the DNA code are that are exons and introns. In the course of this study they have determined that some introns can become extremely large, sometimes over 4.5 million base pairs or a distance of 1.1 kilometers if it was stretched out. Thus a question developes whether the previously described model for splicing works when the two ends of the exon are so far away.
To test this model a messenger RNA was created from existing human genes that contains one of these unique recognition sequences inside the intron. This mRNA was then spliced using an in vitro (in a test tube instead of live cell) system and the results were then examined to determine whether the mRNA product after one splice exist. Then if this is found, to splice this product to see if final product can be produced. |

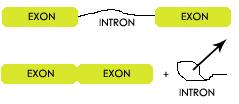 Before messenger RNA (mRNA) messages can be translated into proteins they will undergo additional modifications. One of these modifications is the process of splicing. In this process portions of the mRNA message are removed. Portions that remain in the message to be converted into proteins are called exons. Portions that are removed and later degraded are called introns. Many experiments have shown the widely accepted model of how a splicing takes place. In this model several catalyst identify key codes in the RNA strand. After locating these positions, the catalyst will pull on the intron, reconnect the two exons and disconnect the intron.
Before messenger RNA (mRNA) messages can be translated into proteins they will undergo additional modifications. One of these modifications is the process of splicing. In this process portions of the mRNA message are removed. Portions that remain in the message to be converted into proteins are called exons. Portions that are removed and later degraded are called introns. Many experiments have shown the widely accepted model of how a splicing takes place. In this model several catalyst identify key codes in the RNA strand. After locating these positions, the catalyst will pull on the intron, reconnect the two exons and disconnect the intron. 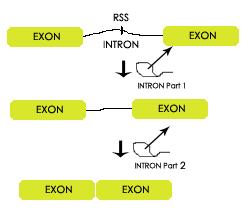 The purpose of this experiment is to provide evidence of a model entitled Recursive Splicing. This model, pictured at right, predicts that an intron will be removed in a multi-step process. The model holds that if recognition sequences are found within the intron then the splicing mechanisms will splice away a portion of the intron then continue to splice the rest of it. This model was previously discovered by a group of scientist working with Drosophila melenagaster (Fruit fly) however the process has not been show to exist in humans.
The purpose of this experiment is to provide evidence of a model entitled Recursive Splicing. This model, pictured at right, predicts that an intron will be removed in a multi-step process. The model holds that if recognition sequences are found within the intron then the splicing mechanisms will splice away a portion of the intron then continue to splice the rest of it. This model was previously discovered by a group of scientist working with Drosophila melenagaster (Fruit fly) however the process has not been show to exist in humans.